Quantitative Proteomics
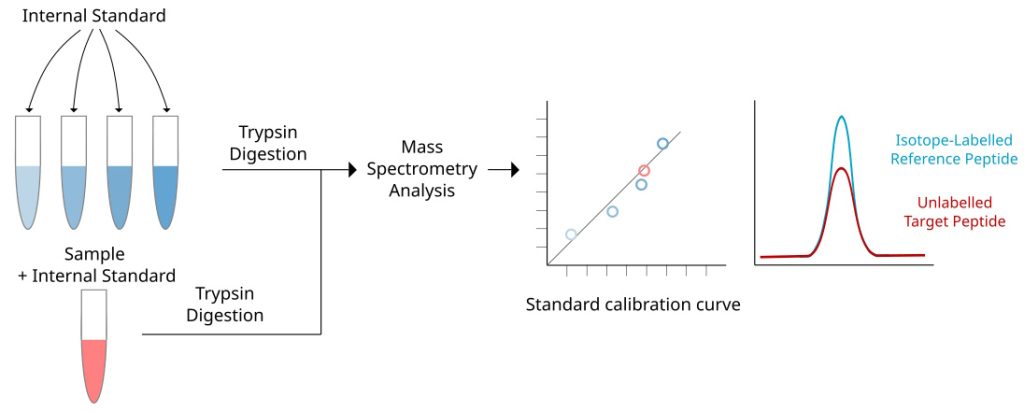
Discovery proteomics enables the identification of thousands of proteins in a sample. However, it provides only limited and unprecise quantitative information about the identified proteins. Quantitative proteomics can advance both discovery and targeted proteomic analyses, facilitating a more in-depth analysis of biological processes.
There are a number of approaches to quantitative proteomics. The highest precision can be reached with heavy isotope labeled proteins, providing direct comparability between the endogenous protein and the reference standard. However, expression and purification of a complete heavy isotope labeled protein is laborious and therefore only reasonable for very small numbers of target proteins. Alternatively, heavy isotope labeled synthetic peptides, short protein segments (ESTs) or synthetic polypeptides (QconCATs) provide a suitable option.
Our method of choice is the QconCAT technology. In contrast to peptides, QconCATs can be spiked at an early step of sample preparation and thus control for all kinds of biases during sample preparation. A well-designed QconCAT enables exact quantification even at an incomplete protein digest. In contrast to full proteins, QconCATs consist of quantitative peptides for up to 50 target proteins and are thus much more cost-effective. It has been shown that QconCATs can reach the same quality as full length labeled proteins (Scott et al, Anal. Chem. 2015, 87, 4429−4435).
Technology
Our proprietary QconCAT technology for absolute protein quantification facilitates the determination of the exact amounts of 10 – 50 of your target proteins with only one QconCAT reference standard in a single read-out.
Advantages
- Accurate quantification
- Full control of sample preparation biases
- Easy multiplexing (quantify a complete proteome using multiple QconCATs)
- Scalable to 1000’s of samples/data points
- Reproducible
- Robust synthesis of large amounts
- Fast assay development
- Compatible with other methods
PolyQuant offers
- Individual QconCATs, tailored for your specific research requirements
- Full service quantification projects
- Fee-for-service quantification
Contact
For more information, assistance and support in planning and execution of your quantitative proteomics project you can contact us:
E-Mail: info[at]polyquant.com
Phone: +49 (0)9405 96999 10
Fax: +49 (0)9405 96999 28



