QconCAT Technology
 QconCATs (Quantification conCATamer) are custom-made synthetic proteins, usually comprising of up to 50 heavy isotope-labelled proteotypic peptides, functioning as true internal standards for absolute protein quantification using mass spectrometry.
QconCATs (Quantification conCATamer) are custom-made synthetic proteins, usually comprising of up to 50 heavy isotope-labelled proteotypic peptides, functioning as true internal standards for absolute protein quantification using mass spectrometry.
Original publications:
- Beynon R. J. et al., Multiplexed absolute quantification in proteomics using artificial QCAT proteins of concatenated signature peptides. Nature Methods Vol. 2 (8), August 2005, 587-589.
- Pratt J. M. et al., Multiplexed absolute quantification for proteomics using concatenated signature peptides encoded by QconCAT genes. Nature Protocols Vol. 1 (2), 2006, 1029-1043.
Download: QconCAT brochure
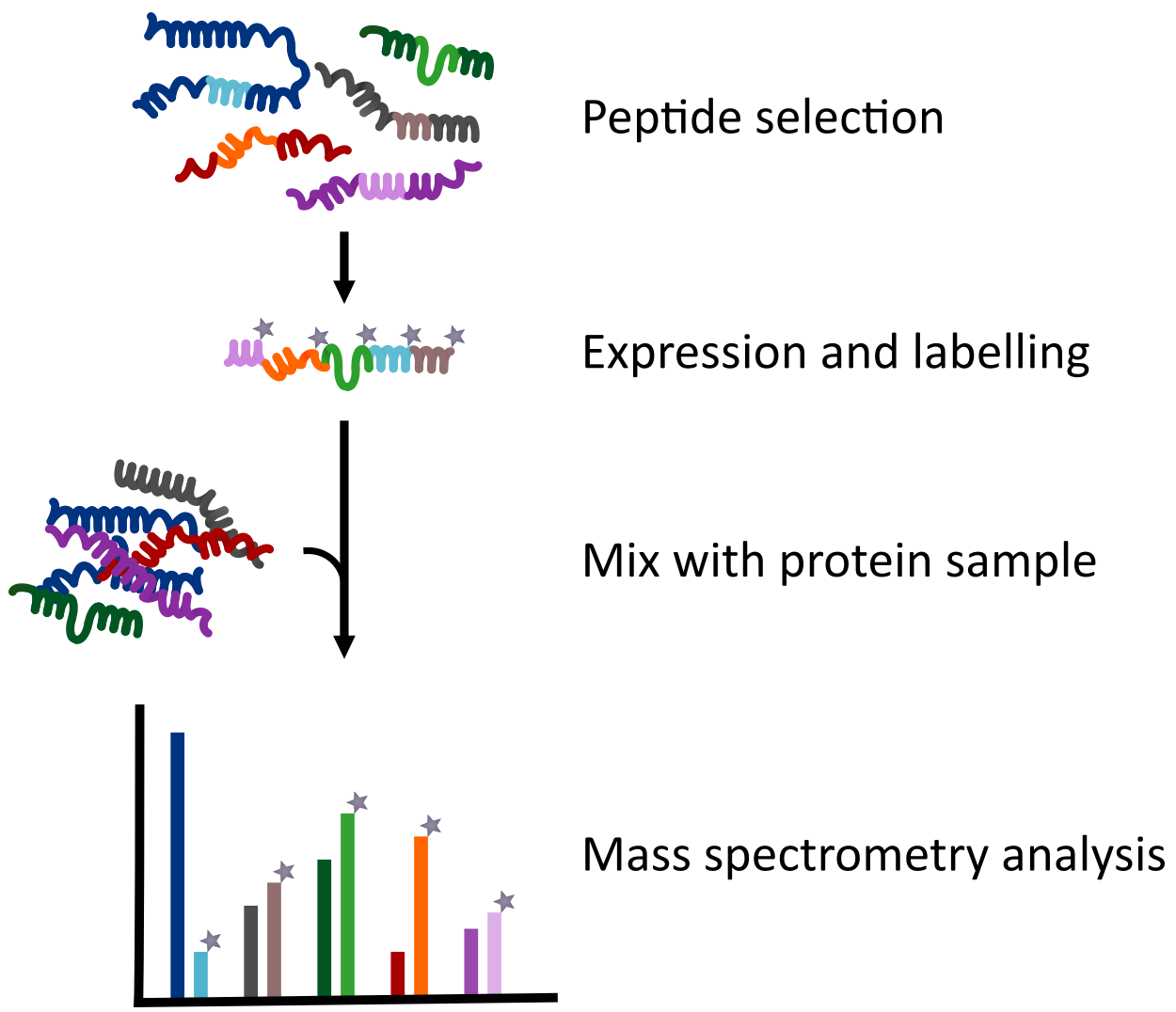
 Proteotypic peptides are selected using either available data or by in silico analysis. The QconCAT is then produced as heavy isotope-labelled synthetic protein.
Proteotypic peptides are selected using either available data or by in silico analysis. The QconCAT is then produced as heavy isotope-labelled synthetic protein. Absolute protein quantification is performed by adding the QconCAT in known quantity to the sample (e. g. a cell extract).
Subsequent proteolysis, for example with trypsin, releases the proteotypic peptides both from the target proteins and the QconCAT for analysis in a mass spectrometer.
The intensity of the reference peptide peaks allows calculation of the exact absolute amount of each target protein.
 QconCATs save time preparing peptide mixes by enabling spiking of the analyte sample with up to 50 reference peptides in a single pipetting step.
QconCATs save time preparing peptide mixes by enabling spiking of the analyte sample with up to 50 reference peptides in a single pipetting step.Besides that, the design of a QconCAT as artificial polypeptide has several other advantages compared to synthetic peptides:
- Synthesize difficult peptides without limitation in peptide length
- Include flanking amino acids to control for variations caused by missed cleavages
- Prevent peptide loss through adsorption to vessel walls
- Release the target peptides in a 1:1 ratio
- Internal control for sample processing by early addition of the QconCAT during sample preparation
 Absolute protein quantification using QconCAT reference standards can be established on all commonly used mass spectrometry platforms. Our technology is compatible with other methods and can be integrated into established quantitative protein analysis workflows.
Absolute protein quantification using QconCAT reference standards can be established on all commonly used mass spectrometry platforms. Our technology is compatible with other methods and can be integrated into established quantitative protein analysis workflows. QconCATs are highly versatile tools for various research projects in academia and industry and have been successfully applied in exploratory studies as well as in high throughput analyses.
A list of selected references using QconCATs by PolyQuant can be found here.

Principal Investigator UMCG, Groningen, The Netherlands | |
|
"Our lab has been applying the QconCATs provided by Polyquant for more than 10 years as they provide a cost-effective way to target complete panels of proteins in combination with the potential to correct for sample preparation losses since the protein standards can be added at the start of the workflow." read more"Our main application is in Systems Medicines-related projects where you require both the accurate quantification of protein levels as well as the information for all proteins in the pathways, which is used to create the computer models. This was for example applied to build personalized models to study the clinical heterogeneity observed in patients with inborn errors of metabolism (PubMed). We are currently extending our panels to study more disease related pathways. Instead of studying protein pathways, we also applied the QconCAT technology to study protein complexes. In these application they will provide a quantitative output of all components in the protein complex and you can also study how these complexes are affected when proteins are absent (PubMed)." |
![]()
![]()
Selected references
- Large Scale Protein Quantitation
- Complex Stoichiometry
- Isoform Specificity and Protein Families
- Signalling Pathways
- Instrument Calibration
- Clinical Proteomics
-
Full humanization of the glycolytic pathway in Saccharomyces cerevisiae
Boonekamp FJ, Knibbe E, Vieira-Lara MA, Wijsman M, Luttik MAH, van Eunen K, Ridder MD, Bron R, Almonacid Suarez AM, van Rijn P, Wolters JC, Pabst M, Daran JM, Bakker BM, Daran-Lapujade P.
Cell Rep. 2022 Jun 28;39(13):111010. [Pubmed]
Boonekamp et al. generated and characterized a yeast strain with a humanized glycolytic pathway, providing a new model to study human glycolysis in a simplified context. In their study they used 6 QconCATs to determine absolute concentrations of glycolytic targets by targeted proteomics. -
Absolute protein quantification of the yeast chaperome under conditions of heat shock
Mackenzie RJ, Lawless C, Holman SW, Lanthaler K, Beynon RJ, Grant CM, Hubbard SJ, Eyers CE.
Proteomics. 2016 Jun 2. [PubMed]
Mackenzie et al. determined copy per cell values for 49 key chaperones in S. cerevisiae under conditions of normal growth and heat shock, by selected reaction monitoring (SRM) mass spectrometry using QconCATs as reference peptides. - Direct and Absolute Quantification of over 1800 Yeast Proteins via Selected Reaction Monitoring
Lawless C, Holman SW, Brownridge P, Lanthaler K, Harman VM, Watkins R, Hammond DE, Miller RL, Sims PF, Grant CM, Eyers CE, Beynon RJ, Hubbard SJ.
Mol Cell Proteomics. 2016 Apr;15(4):1309-22. [PubMed]
Lawless et al. quantified over 1800 S. cerevisiae proteins by selected reaction monitoring (SRM) mass spectrometry using over 100 QconCATs.
-
CRAF dimerization with ARAF regulates KRAS-driven tumor growth
Venkatanarayan A, Liang J, Yen I, Shanahan F, Haley B, Phu L, Verschueren E, Hinkle TB, Kan D, Segal E, Long JE, Lima T, Liau NPD, Sudhamsu J, Li J, Klijn C, Piskol R, Junttila MR, Shaw AS, Merchant M, Chang MT, Kirkpatrick DS, Malek S.
Cell Rep. 2022 Feb 8;38(6):110351. [PubMed]
Venkatanarayan et al. used quantitative proteomics to demonstrate increased levels of CRAF:ARAF dimers in KRAS mutant cells and PRM analysis to quantify the stoichiometric relationships between wt and mutant MAPK components using a QconCAT incorporating wild type and mutant sequences for detection of disease associated mutations. -
PIKES Analysis Reveals Response to Degraders and Key Regulatory Mechanisms of the CRL4 Network
Reichermeier KM, Straube R, Reitsma JM, Sweredoski MJ, Rose CM, Moradian A, den Besten W, Hinkle T, Verschueren E, Petzold G, Thomä NH, Wertz IE, Deshaies RJ, Kirkpatrick DS.
Mol Cell. 2020 Jan 15. [Pubmed]
Reichermeier et al. determined the stoichiometric relationships between approx. 30 proteins in cell lysates and immunoprecipitated samples using targeted proteomics and a QconCAT comprising of 67 peptides for 31 proteins. -
Stoichiometry, Absolute Abundance, and Localization of Proteins in the Bacillus cereus Spore Coat Insoluble Fraction Determined Using a QconCAT Approach
Stelder SK, Benito de Moya C, Hoefsloot HCJ, de Koning LJ, Brul S, de Koster CG.
J Proteome Res. 2018 Feb 2;17(2):903-917 [Pubmed]
Stelder et al. quantified crucial spore proteins using a QconCAT reference standard, determining the absolute abundance of 21 proteins, covering approx. 5.66% of the total spore weight in wild type and approx. 4.13% in the ΔCotE mutant. -
Rigorous determination of the stoichiometry of protein phosphorylation using mass spectrometry
Johnson H, Eyers CE, Eyers PA, Beynon RJ, Gaskell SJ.
Journal of the American Society for Mass Spectrometry 2009 Dec;20(12):2211-20.
[PubMed]
Johnson et al. use QconCATs to determine absolute protein concentrations and the stoichiometry of phosphorylation at predefined sites.
Loss of hepatic SMLR1 causes hepatosteatosis and protects against atherosclerosis due to decreased hepatic VLDL secretion
van Zwol W, Rimbert A, Wolters JC, Smit M, Bloks VW, Kloosterhuis NJ, Huijkman N, Koster MH, Tharehalli U, de Neck SM, Bournez C, Fuh MM, Kuipers J, Rajan S, de Bruin A, Ginsberg HN, van Westen GJP, Hussain MM, Scheja L, Heeren J, Zimmerman P, van de Sluis B, Kuivenhoven JA.
Hepatology. 2023 Nov 1. [Pubmed]
Van Zwol et al. identified small leucine-rich protein 1 (SLMR1) as a player in the VLDL biogenesis pathway and assessed its role by generating a liver-specific knockout mouse. In their study they used a targeted proteomics approach to measure protein levels of 11 apolipoproteins using a mix of QconCATs and isotopically labeled standard peptides.Characterization of CYP2B6 K262R allelic variants by quantitative allele-specific proteomics using a QconCAT standard
Barber J, Russell MR, Rostami-Hodjegan A, Achour B.
J Pharm Biomed Anal. 2020 Jan 30;178:112901. [Pubmed]
Barber et al. used a QconCAT standard for targeted proteomics, simultaneously determining protein abundance and missense polymorphisms.-
COMMD Family Regulates Plasma LDL Levels and Attenuates Atherosclerosis Through Stabilizing the CCC Complex in Endosomal LDLR Trafficking
Fedoseienko A, Wijers M, Wolters JC, Dekker D, Smit M, Huijkman N, Kloosterhuis N, Klug H, Schepers A, Willems van Dijk K, Levels JH, Billadeau DD, Hofker MH, van Deursen J, Westerterp M, Burstein E, Kuivenhoven JA, van de Sluis B.
Circ Res. 2018 Mar 15. [Pubmed]
Fedoseienko et al. used QconCATs to quantify the protein concentrations of the COMMDs, components of retromer, the CCC and WASH complexes, etc in the samples of the different mouse models.
-
A Kinase Interacting Protein 1 regulates mitochondrial protein levels in energy metabolism and promotes mitochondrial turnover after exercise
Kirsten T. Nijholt, Pablo I. Sánchez-Aguilera, Belend Mahmoud, Albert Gerding, Justina C. Wolters, Anouk H. G. Wolters, Ben N. G. Giepmans, Herman H. W. Silljé, Rudolf A. de Boer, Barbara M. Bakker, B. Daan Westenbrink
Sci Rep. 2023; 13: 18822. Published online 2023 Nov 1. [Pubmed]
Nijhoult et al., studied the influence of A Kinase Interacting Protein 1 (AKIP1) on mitochondrial function and adaptation in response to exercise in vivo. They could show that levels of proteins in mitochondrial energy metabolism and related pathways was changed in hearts from mice with cardiomyocyte-specific overexpression of AKIP1 by performing quantitative proteomics with QconCATs from PolyQuant targeting 38 proteins involved in fatty acid β-oxidation, tricarboxylic acid cycle (TCA), oxidative phosphorylation (OXPHOS), substrate transport and antioxidant activity. -
Species-specific metabolic reprogramming in human and mouse microglia during inflammatory pathway induction
Sabogal-Guáqueta AM, Marmolejo-Garza A, Trombetta-Lima M, Oun A, Hunneman J, Chen T, Koistinaho J, Lehtonen S, Kortholt A, Wolters JC, Bakker BM, Eggen BJL, Boddeke E, Dolga A.
Nat Commun. 2023 Oct 13;14(1):6454. [Pubmed]
Sabogal-Guáqueta et al., investigated the effects of lipopolysaccharides (LPS) on mouse microglia and human microglia-like cells at the protein level, performing both label-free and targeted proteomics. They quantitatively determined expression levels of 11 TCA cycle enzymes, using QconCAT reference standards. The targeted approach enabled them to observe an increased abundance of the isoforms PFKM and PFKP in human but not mouse microglia. -
Effects of lysine deacetylase inhibitor treatment on LPS responses of alveolar-like macrophages
Russo S, Kwiatkowski M, Wolters JC, Gerding A, Hermans J, Govorukhina N, Bischoff R, Melgert BN.
J Leukoc Biol. 2023 Oct 9:qiad121. [Pubmed]
Russo et al., investigated if the anti-inflammatory effect of lysine deacetylase inhibitors correlated with metabolic changes in macrophages. Performing targeted proteomic analysis of 59 proteins (encoded on QconCATs from PolyQuant) involved in the major metabolic pathways revealed no significant alterations of their proteins levels. However, their data indicates that protein ubiquitination may be the driver of the anti-inflammatory effects of lysine deacetylase inhibitors. -
Absolute proteomic quantification reveals design principles of sperm flagellar chemosensation
Trötschel C, Hamzeh H, Alvarez L, Pascal R, Lavryk F, Bönigk W, Körschen HG, Müller A, Poetsch A, Rennhack A, Gui L, Nicastro D, Strünker T, Seifert R, Kaupp UB.
EMBO J. 2020 Feb 17;39(4) [PubMed]
In this study, Trötschel et al. used a QconCAT from PolyQuant to absolutely quantify 19 proteins of a flagellar chemosensory signalling pathway from sperm of the sea urchin Arbacia punctulata.
-
QCAL--a novel standard for assessing instrument conditions for proteome analysis
Eyers CE, Simpson DM, Wong SC, Beynon RJ, Gaskell SJ.
J Am Soc Mass Spectrom. 2008 Sep;19(9):1275-80. [Pubmed]
Original publication introducing the QCAL calibration standard for mass spectrometry instrumentations. -
RePLiCal: A QconCAT Protein for Retention Time Standardization in Proteomics Studies
Holman SW, McLean L, Eyers CE.
J Proteome Res. 2016 Mar 4;15(3):1090-102 [Pubmed]
Original publication introducing the RePLiCal retention time standard. -
Platform Independent Protein-Based Cell-Of-Origin Subtyping of Diffuse Large B-cell Lymphoma in Formalin-Fixed Paraffin-Embedded Tissue
Reinders J, Altenbuchinger M, Limm K, Schwarzfischer P, Scheidt T, Strasser L, Richter J, Szczepanowski M, Huber CG, Klapper W, Spang R, Oefner PJ.
Sci Rep. 2020 May 12;10(1):7876. [Pubmed]
In their study, Reinders et al. used RePLiCal for normalization of retention times by linear fitting.
-
Quantification of drug metabolising enzymes and transporter proteins in the paediatric duodenum via LC-MS/MS proteomics using a QconCAT technique
Goelen J, Farrell G, McGeehan J, Titman CM, J W Rattray N, Johnson TN, Horniblow RD, Batchelor HK.
Eur J Pharm Biopharm. 2023 Aug 23:S0939-6411(23)00214-X. [Pubmed]
Goelen et al. used a QconCAT to simultaneously quantify 21 proteins of the three DMET-protein families (transporters, CYP and UGT-enzymes) using pinch biopsies from the paediatric duodenum. -
Proteomic quantification of receptor tyrosine kinases involved in the development and progression of colorectal cancer liver metastasis.
Vasilogianni AM, Al-Majdoub ZM, Achour B, Peters SA, Rostami-Hodjegan A, Barber J.
Front Oncol. 2023 Feb 20;13:1010563. [Pubmed]
Vasilogianni et al. assessed protein abundance of 21 receptor tyrosine kinases (RTKs) in 15 healthy and 18 cancerous liver samples using a QconCAT to validate protein abundance. Their work allowed identification of proteins with reduced protein levels in cancerous liver samples, but also proteins whose protein levels were increased. -
Personalised modelling of clinical heterogeneity between medium-chain acyl-CoA dehydrogenase patients
Odendaal C, Jager EA, Martines AMF, Vieira-Lara MA, Huijkman NCA, Kiyuna LA, Gerding A, Wolters JC, Heiner-Fokkema R, van Eunen K, Derks TGJ, Bakker BM
BMC Biol. 2023 Sep 4;21(1):184. [Pubmed]
Odendaal et al., built and validated a kinetic model of the human liver mitochondrial fatty acid oxidation (mFAO). They used QconCATs from PolyQuant to determine absolute quantities of 9 key proteins, observing a clear distinct pattern of the SCAD protein in the asymptomatic patient. Their data underlines that kinetic models are powerful tools, complementing models based on genomic data. -
Cov2MS: An Automated and Quantitative Matrix-Independent Assay for Mass Spectrometric Measurement of SARS-CoV-2 Nucleocapsid Protein
Van Puyvelde B, Van Uytfanghe K, Van Oudenhove L, Gabriels R, Van Royen T, Matthys A, Razavi M, Yip R, Pearson T, Drouin N, Claereboudt J, Foley D, Wardle R, Wyndham K, Hankemeier T, Jones D, Saelens X, Martens G, Stove CP, Deforce D, Martens L, Vissers JPC, Anderson NL, Dhaenens M.
Anal Chem. 2022 Dec 20;94(50):17379-17387. [Pubmed]
Van Puyvelde et al. improved Cov-MS by adding SISCAPA technology to enrich proteotypic peptides of the SARS-CoV-2 nucleocapsid (N) protein from trypsin-digested patient samples. The Cov2MS assay is compatible with most matrices including nasopharyngeal swabs, saliva, and plasma, has increased sensitivity and a strong positive correlation with qPCR detection beyond a quantification cycle of 30-31. -
A fast and sensitive absolute quantification assay for the detection of SARS-CoV-2 peptides using parallel reaction monitoring mass spectrometry
Gajbhiye A, Nalbanta A, Heunisa T, Sidgwick F, Porter A, Tahab Y, Trost M
Journal of Proteomics. Vol. 265, 15 August 2022. [Pubmed]
Gajbhiyea et al. developed a high-throughput PRM-MS assay to directly detect viral peptides in nasopharyngeal swab samples. The assay enables sensitive detection and absolute quantification of SARS-CoV-2 nucleocapsid peptides with short turn-around times by using the Cov-MS isotopically labelled synthetic polypeptide as reference standard. -
Proteomics of colorectal cancer liver metastasis: A quantitative focus on drug elimination and pharmacodynamics effects
Vasilogianni AM, Al-Majdoub ZM, Achour B, Peters SA, Rostami-Hodjegan A, Barber J.
Br J Clin Pharmacol. 2022 Feb;88(4):1811-1823. [Pubmed]
Vasilogianni et al. used QconCATs to quantify drug-metabolising enzymes, transporters, receptor tyrosine kinases (RTK) and protein markers in samples from colorectal cancer liver metastasis (CLRM) patients. The detected alterations in protein abundance may provide valuable information for diagnosis and therapeutic intervention. -
Quantitative Proteomics of Hepatic Drug-Metabolizing Enzymes and Transporters in Patients with Colorectal Cancer Metastasis
Vasilogianni AM, Al-Majdoub ZM, Achour B, Peters SA, Barber J, Rostami-Hodjegan A.
Clin Pharmacol Ther. 2022 May 3. [Pubmed]
Vasilogianni et al. studied the impact of liver cancer metastasis on protein abundance of 22 drug-metabolizing enzymes (DMEs) and 25 transporters. They used targeted proteomics and QconCATs as reference standards for analysis of microsome preparations from individuals and cancer patients. -
A family of QconCATs (Quantification conCATemers) for the quantification of human pharmacological target proteins
Vasilogianni AM, El-Khateeb E, Achour B, Alrubia S, Rostami-Hodjegan A, Barber J, Al-Majdoub ZM.
J Proteomics. 2022 Jun 15;261:104572. [Pubmed]
In this report, the authors describe the development and characterization of two QconCAT constructs for quantification of 24 enzymes and 21 receptor tyrosine kinases (RTKs), complementing two previously reported QconCATs for the quantification of key enzymes and drug transporters. The QconCATs were successfully applied in quantification of target proteins in human liver. -
Targeted LC-MS/MS for the evaluation of proteomics biomarkers in the blood of neonates with necrotizing enterocolitis and late-onset sepsis
Chatziioannou AC, Wolters JC, Sarafidis K, Thomaidou A, Agakidis C, Govorukhina N, Kuivenhoven JA, Bischoff R, Theodoridis G.
Anal Bioanal Chem. 2018 Nov;410(27):7163-7175. [Pubmed]
Chatziioannou et al. used QconCATs and synthetic peptides belonging to 47 protein markers for a prospective case-control study evaluating serum proteomics profiles. They were able to define two panels of three proteins each that allow highly sensitive diagnosis of late-onset sepsis (LOS) and differential diagnosis between LOS and necrotizing enterocolitis. -
Non-uniformity of Changes in Drug-Metabolizing Enzymes and Transporters in Liver Cirrhosis: Implications for Drug Dosage Adjustment
El-Khateeb E, Achour B, Al-Majdoub ZM, Barber J, Rostami-Hodjegan A.
Mol Pharm. 2021 Sep 6;18(9):3563-3577.[Pubmed]
El-Katheeb et al. used QconCAT-based targeted proteomics to determine the absolute abundance of 51 drug-metabolizing enzymes and transporters in human liver microsomes across the three degrees for cirrhosis severity to determine their impact on the predictive performance of PBPK (physiologically based pharmacokinetic) models. Their work demonstrates the utility of proteomics-informed PBPK modeling for drug-specific dose adjustments in liver cirrhosis. -
Cov-MS: A Community-Based Template Assay for Mass-Spectrometry-Based Protein Detection in SARS-CoV-2 Patients
Bart Van Puyvelde, Katleen Van Uytfanghe, Olivier Tytgat, Laurence Van Oudenhove, Ralf Gabriels, Robbin Bouwmeester, Simon Daled, Tim Van Den Bossche, Pathmanaban Ramasamy, Sigrid Verhelst, Laura De Clerck, Laura Corveleyn, Sander Willems, Nathan Debunne, Evelien Wynendaele, Bart De Spiegeleer, Peter Judak, Kris Roels, Laurie De Wilde, Peter Van Eenoo, Tim Reyns, Marc Cherlet, Emmie Dumont, Griet Debyser, Ruben t’Kindt, Koen Sandra, Surya Gupta, Nicolas Drouin, Amy Harms, Thomas Hankemeier, Donald J. L. Jones, Pankaj Gupta, Dan Lane, Catherine S. Lane, Said El Ouadi, Jean-Baptiste Vincendet, Nick Morrice, Stuart Oehrle, Nikunj Tanna, Steve Silvester, Sally Hannam, Florian C. Sigloch, Andrea Bhangu-Uhlmann, Jan Claereboudt, N. Leigh Anderson, Morteza Razavi, Sven Degroeve, Lize Cuypers, Christophe Stove, Katrien Lagrou, Geert A. Martens, Dieter Deforce, Lennart Martens, Johannes P. C. Vissers, and Maarten Dhaenens.
JACS Au 2021, 1, 6, 750–765.[Pubmed]
The authors established a consortium consisting of 15 academic laboratories and several industrial partners to identify peptides for MS-based detection of SARS-CoV-2 infection. They describe the full pipeline for developing a fast and sensitive MS-based assay for detection of viral presence directly in clinical samples using conventional instrumentation.



