Metaproteomics
- Determine the biodiversity of microbial or viral communities and multi-organism systems
- Identify and quantify proteins from microbial communities
- Analyze the expressed proteins for each organism
- Quantification of per species biomass
- Track substrate metabolisms
Metaproteomics is a rapidly growing research area, addressing the microbial composition of environmental samples (soil, feces, sewage, water etc) [further reading].
We have developed metaproteomics workflows covering protein extraction, peptide separation, LC-MS/MS measurements and data analysis. Our techniques allow us to determine the organisms present in a sample and their proportions as well as comprehensive analysis of the expressed proteins for each organism.

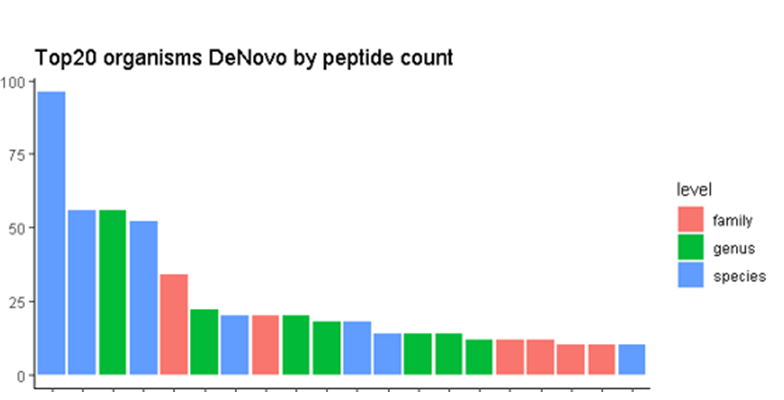
Example:
A protein sample of sewage sludge of unknown composition was solubilized and 30 µg of protein were digested with trypsin. The peptides were then fractionated by binding the peptides to a filter, followed by elution with increasing concentrations of acetonitrile (ACN). The samples were measured on an Orbitrap HF-X mass spectrometer coupled to an AQUITY UPLC system, using a 25 cm M‑Class column. Gradients of 60, 130 or 300 min were used to separate the peptides. Peptides were identified using a de novo peptide sequencing algorithm and by a sectioning database approach, searching against the whole Uniprot bacteria database. The organisms identified in the first round of analysis were used to construct specific databases for more in-depth analysis of the provided sample.
Our offer
We offer established workflows and individual support for research projects and routine applications including:
Various sample preparation techniques and LC-MS/MS workflows
Automated and manually curated data analysis workflows
You receive
Comprehensive analysis, performed and evaluated by experienced professionals
Contact
For more information, assistance and support or to ask for a quote, please contact us:
E-Mail: info[at]polyquant.com
Phone: +49 (0)9405 96999 10
Fax: +49 (0)9405 96999 28



