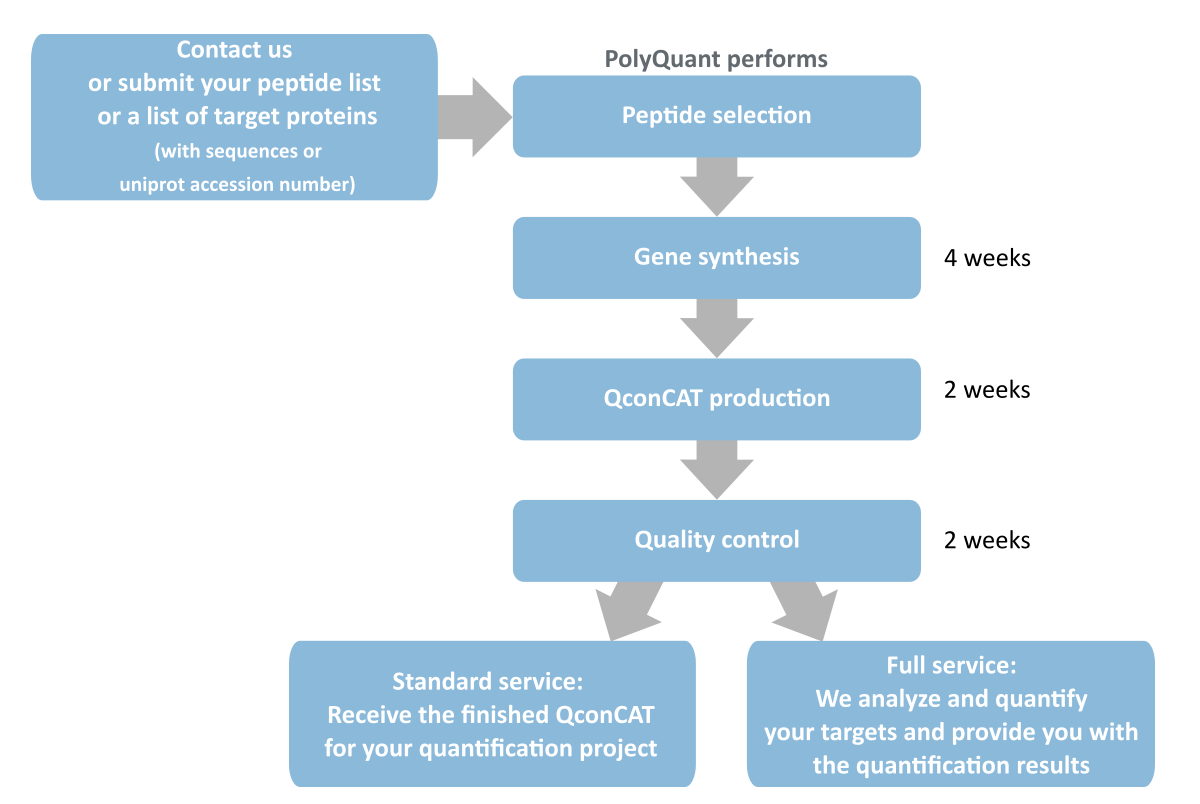
QconCAT – absolute protein quantification

PolyQuant’s proprietary QconCAT technology for absolute protein quantification allows you to:
- Absolutely quantify several tens of proteins with a single QconCAT
- Quantify whole proteomes in a single experiment using several QconCATs
- Receive large amounts of QconCAT for high throughput analyses
- Compare data between laboratories and instruments
- Minimize sample preparation bias by early addition of the internal standard
more information
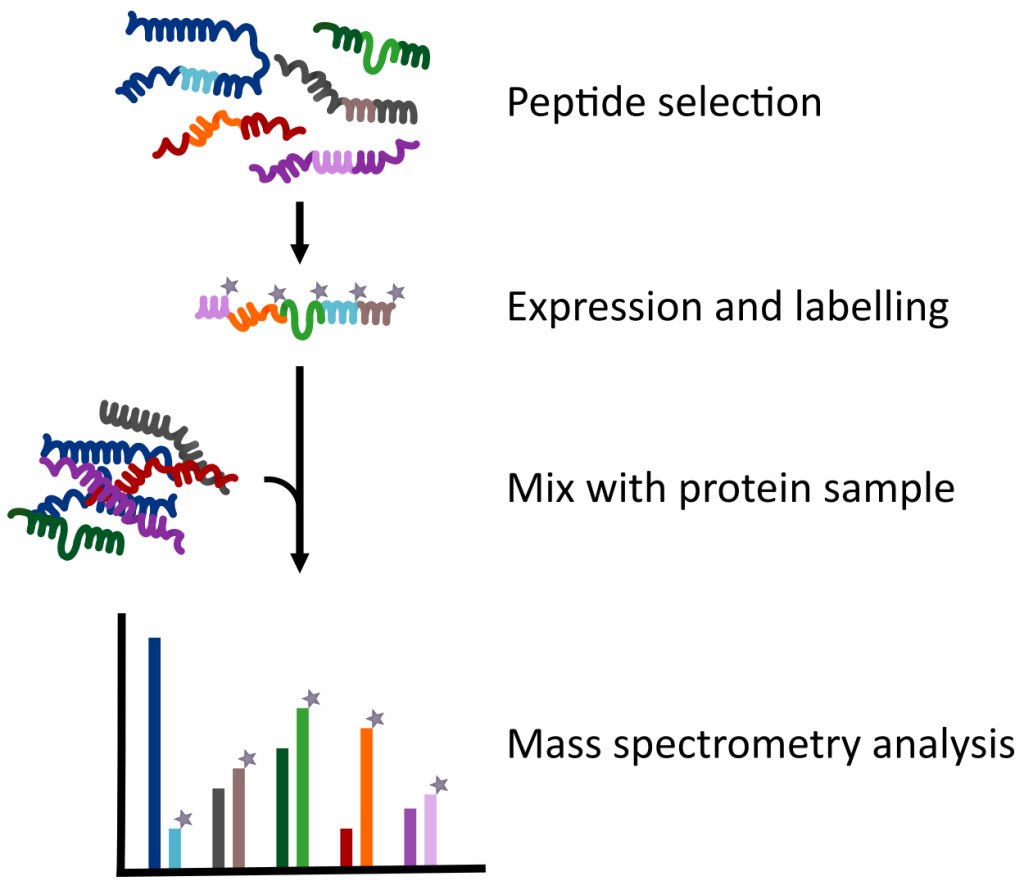
QconCATs (Quantification conCATamer) are custom-made synthetic proteins, usually comprising of up to
50 heavy isotope-labelled proteotypic peptides, functioning as true internal standards for protein quantification using mass spectrometry.
- Skip time-consuming preparation of peptide mixes and spike your analyte sample with up to 50 reference peptides in a single pipetting step.
- Receive an individually produced isotope-labelled QconCAT protein, based on a detailed analysis of your requirements.
- Our experienced team provides support from selection of proteotypic peptides (based on available data or by in silico analysis) to setting up your targeted method in Skyline (receive peptide libraries and iRT values).
Our offer
Custom-made QconCAT reference standards are individually produced for each project as heavy-labelled protein with the label of your choice.
Each QconCAT is quality controlled via SDS-PAGE (purity) and LC-MS/MS (identity, labelling efficiency, quantity).
Delivery costs depend on destination and packaging. Bioinformatic support for data analysis is available upon request.
For more information and support you can contact us:
E-Mail: info[at]polyquant.com
Phone: +49 (0)9405 96999 10
Fax: +49 (0)9405 96999 28
Delivery costs depend on destination and packaging. Bioinformatic support for data analysis is available upon request.
For more information and support you can contact us:
E-Mail: info[at]polyquant.com
Phone: +49 (0)9405 96999 10
Fax: +49 (0)9405 96999 28
 Benefit from our profound expertise in dealing with comprehensive protein quantification projects.
Benefit from our profound expertise in dealing with comprehensive protein quantification projects.Receive support and assistance at every step of the project, from protocol development, peptide selection to measurement, data analysis and comprehensive reporting.
For more information, assistance and support in planning and execution of your quantitative proteomics project you can contact us:
E-Mail: info[at]polyquant.com
Phone: +49 (0)9405 96999 10
Fax: +49 (0)9405 96999 28